Breaking news – by Prof Bob Lightowlers
Professor Dame Sally Davies, the Governments Chief Medical Officer, has today given support to a change in the law that will allow the form of mitochondrial replacement therapy that is being pioneered here in Newcastle, has we mentioned before.
Its great news for any prospective mother with mitochondrial DNA disease who is concerned about transmitting the disease to their children.
Most media is covering this – BBC, Guardian, Telegraph – but if you are still confused, we thought it would be good to recap the essential points:

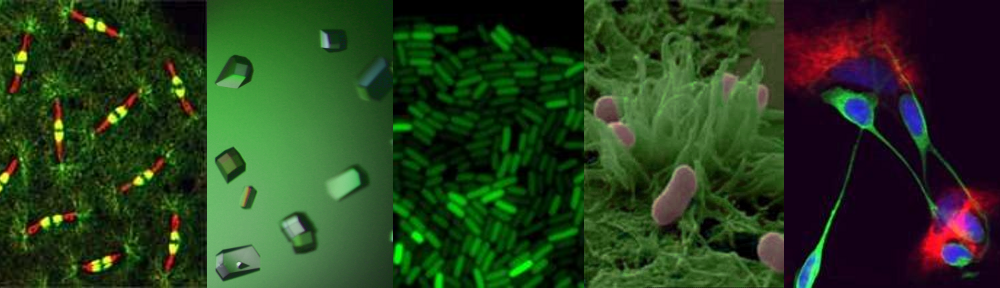
Electron micrograph of a cell (coloured blue) revealing part of the mitochondrial structure (orange) within. The entire length of the mitochondrion is about 5 micrometres.
– Mitochondria are crucial structures found in all cells in our body and they take our common foodstuffs such as fats and sugars and turn them into energy. They have their own genetic element, mitochondrial (mt) DNA. Much smaller than our chromosomes, mtDNA is essential for energy production.
– In 1988, scientists in the UK and US recognised that certain diseases were caused by mutations in mtDNA with the main disorders related to your muscle tissue and the brain.
– It is estimated that at least 1:10,000 people suffer from disorders associated with defects in mtDNA – that’s more than 6,000 people in the UK.
– Mitochondrial DNA is only transmitted to babies by their mothers. Unfortunately, as you inherit your mothers mitochondria, diseases caused by mtDNA mutations are inadvertently transmitted from the mother.
– Doug Turnbull, Director of the Wellcome Trust Centre for Mitochondrial Research (WTCMR) and I have wondered since the 1990s whether it would be possible to prevent the transmission of the faulty mtDNA from the mothers to their children by transferring the nucleous from an egg of the mother carrying these mtDNA mutations into a healthy egg whose nucleous had been removed, and then fertilise it
– Professor Mary Herbert, Doug and a group of us from the Mitochondrial Research Group were able to show that such a swap could be performed without any or very low levels of the defective mtDNA being transferred. Importantly, there was also no defect detectable in the reconstituted cells.
– In August 2012, the government asked the Human Fertility and Embryological Authority (HFEA) to find out what the general public thought of the procedure. The results were collated and the Human Fertility and Embryological Authority made a recommendation to Government. There was an overall support for the new technology with only 10% being fairly or strongly against the concept of mitochondrial gene replacement
To understand all the details and find out the whole story, see my post a while ago here.
The Wellcome Trust also has a very nice video explaining the science behind the news story:
Links
Wellcome Trust Centre for Mitochondrial Research http://www.newcastle-mitochondria.com/
Human Fertility and Embryological Authority (HFEA) http://www.hfea.gov.uk/index.html
HFEA mitochondria puclib consultation 2012 http://www.hfea.gov.uk/6896.html