On a bright, sunny day in Newcastle, Soapbox Science took place on 17th June for the third time in Newcastle’s Monument, supported by the Newcastle University Faculty of Medical Sciences Engagement Team, and by the organising team from the Institute for Cell and Molecular Biosciences.
During the event, Grey Street was converted into an area for lively discussion of scientific subjects between researchers and the general public. Inspired by Hyde Park Corner‘s political debates, our scientists stood on soapboxes wearing lab coats and communicated their work to the passing crowds. There were no microphones, PowerPoint presentations, or posters, only materials from everyday life to support the scientific talks.
Two researchers from the Institute for Cell and Molecular Biosciences, Professor Janet Quinn and postgraduate student Clare Willis, had been selected for a speaker’s role in the event, along with a number of other researchers in diverse scientific fields. Prof. Janet Quinn and her topic “Killer Fungus” grabbed the attention of many younger onlookers by creating Candida, Schizosaccharomyces and other representatives of the fungal Kingdom from colourful Play-Doh in Petri dishes. Clare Willis demonstrated the possibility for survival of a living organism for thousands of years as part of her topic: “Spores: the ultimate survivors”. An air balloon represented the bacterial cell as a target for antibiotics, easily destroyed by a pin, with an unbreakable bacterial spore shown as a much more resilient tennis ball.
A number of volunteers from the Institute for Cell and Molecular Biosciences actively participated in the event. Here, we’ve compiled comments from a few of those who contributed to making the day a success.

Yana performing her duties as part of the evaluation team, gathering data from Soapbox visitors
Yana Wade, researcher in the Centre for Bacterial Cell Biology and Soapbox Science volunteer:
“During my stay at Newcastle University, the secondment host for my Marie Curie Individual Fellowship, I had the fantastic opportunity to be involved in the Soapbox Science festival both as a public engagement volunteer and as part of the evaluation team. As an active researcher I understand the importance of communicating with the public to highlight exactly what scientists do behind the doors of the laboratory, and how they tackle issues that are having a major impact in our daily lives.
The challenges given to the speaker’s at this event were: as an outstanding scientist performing cutting edge research, would it be possible to explain the complex scientific topics to a non-expert audience?Would it be realistic to grab the attention of the general public out shopping on a busy central street who are normally put-off by the often challenging ideas presented by science? Could the scientists explain their research that could capture the imagination of primary school children?
Nevertheless, the presenters at Soapbox Science successfully achieved these goals. Instead of a purely academic presentation of their work, they shared their passion about science by demonstrating how exciting, interesting and important science is in solving immediate and longer term problems faced by society. The event also provided an opportunity for the public to interact directly with scientists in an informal and friendly setting.
The other fascinating aspect of the event was the aim to promote women in science. The female speakers, comprising different academic levels from student to professor, demonstrated their accomplishments in state-of-the-art research, and represented an example of a successful career path for budding scientists in academia. These brilliant girls are addressing the gender-related stereotypes in science.

Our Soapbox speakers were delighted with the public response
By participating in this event as a volunteer, it gave me the means to contribute to bringing science to society in an open and understandable way. I was able to question people who took part in the event and gather feedback that will help evaluate the impact and perception of the topics covered and how they think the subjects could help in shaping society, but also, more generally, to improve the event in the future.
Given my experience, I would like to encourage people to support the Soapbox Science by actively participating in the future.”
Rashmi Bhardwaj-Gosling, PhD student in the Institute for Health & Society, and Soapbox speaker:

Rashmi speaks from her Soapbox
“I had a great afternoon speaking at the event. Speaking to the general public was such a novel and refreshing approach for me in comparison to speaking to my usual audience! Soapbox Science is such a great idea where instead of waiting for the public to come to us, we got out there and took our science to the streets. The audience was very engaging and genuinely interested in finding out about my research work on urinary catheters and what can be done to avoid the catheter related infections. It was brilliantly organised event and weather certainly helped to captivate the crowd. So well done to everyone who took part and in organising the event.”
Clare Willis, researcher in CBCB and Soapbox speaker:
“Soapbox Science was a fantastic event. As a speaker I really enjoyed the challenge of having to explain my research in simple easy-to-understand ways, making it accessible, interesting and exciting to a public audience. I got some great questions and I think my talk really got the public thinking. I feel I’ve made Newcastle a little more knowledgeable about my topic, bacterial spores!”

Clare on her Soapbox, presenting her science to the public
Katherine Jacques and Helen Atkinson, Faculty of Medical Sciences Engagement Team and Soapbox organisers:
“We really enjoyed organising Soapbox Science this year and had a great time on the day. The enthusiasm from the speakers and volunteers to engage with the public was fantastic and it was wonderful to see so many people stopping to listen and ask questions.
We would like to say a big thank you to the speakers, volunteers and our fellow organisers for their contribution and making the day a huge success. We’re already looking forward to another successful Soapbox Science event for next year!”
You can also read a blog post from one of this year’s speakers, Luiza Andrade, on MicrobeBlog from the Society for Applied Microbiology here.
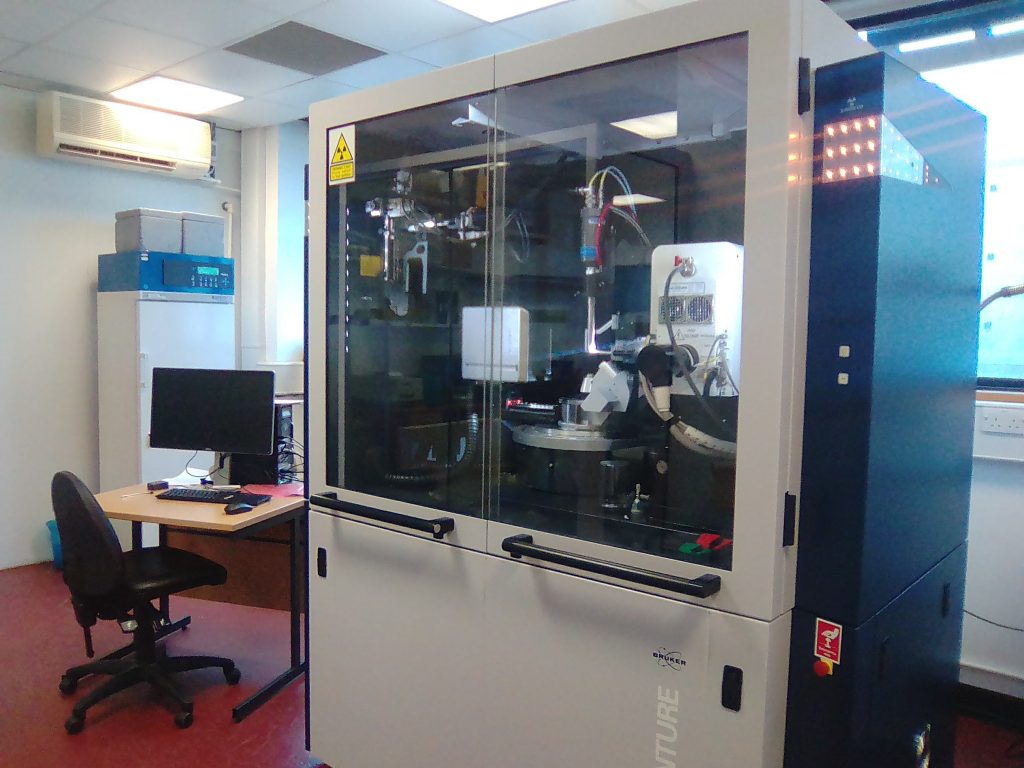
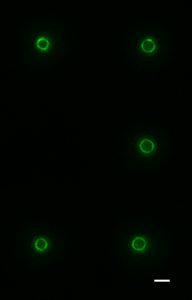
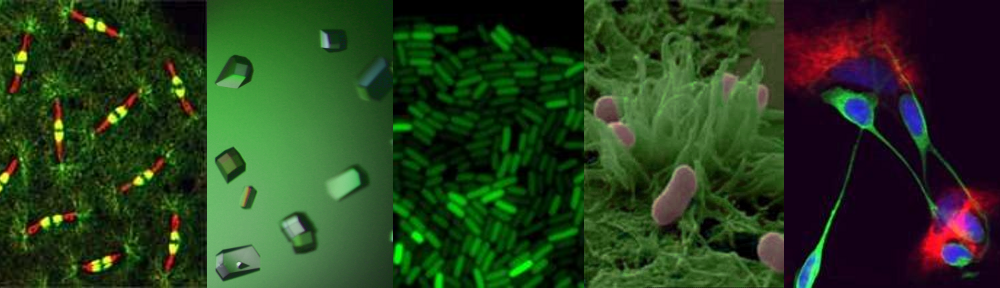







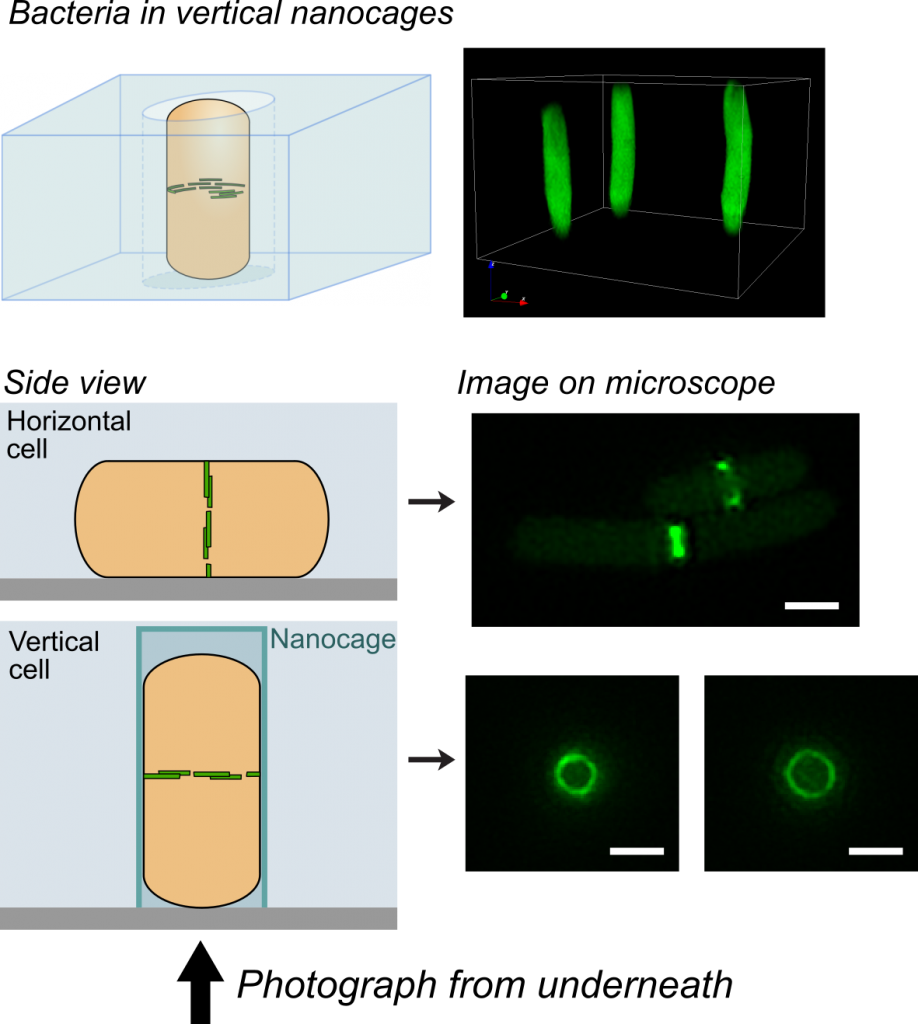
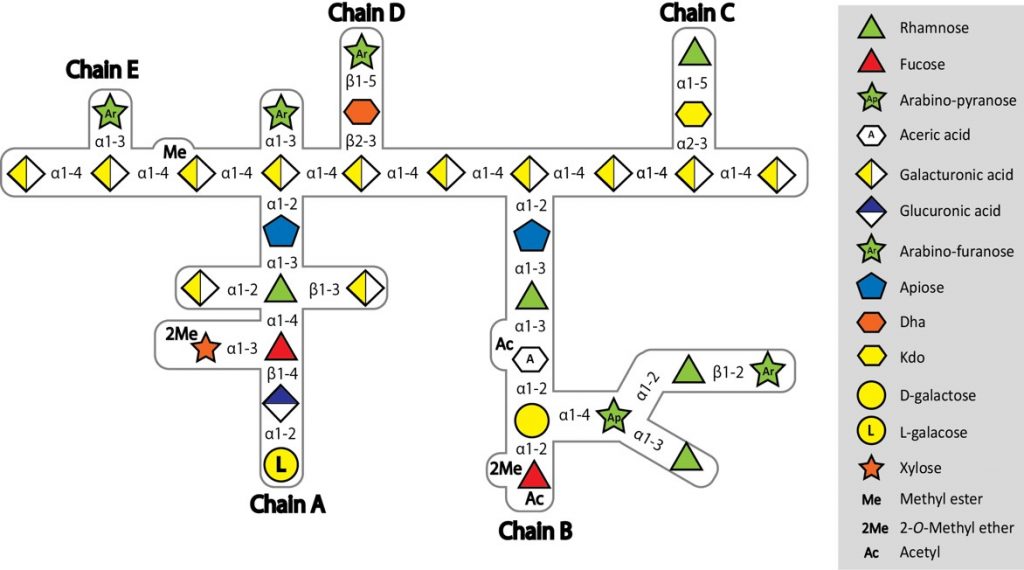






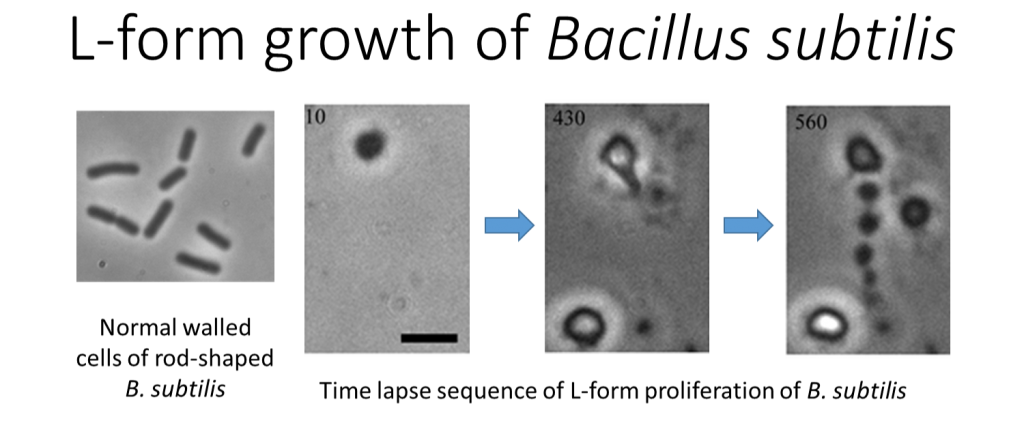


 It is sadly not unusual for frail patients to contract serious infections in hospital. Clinicians are of course ready for this and would have responded by administering powerful antibiotics, hoping to kill the invading bacteria. Unfortunately, increasingly in the modern era, the
It is sadly not unusual for frail patients to contract serious infections in hospital. Clinicians are of course ready for this and would have responded by administering powerful antibiotics, hoping to kill the invading bacteria. Unfortunately, increasingly in the modern era, the 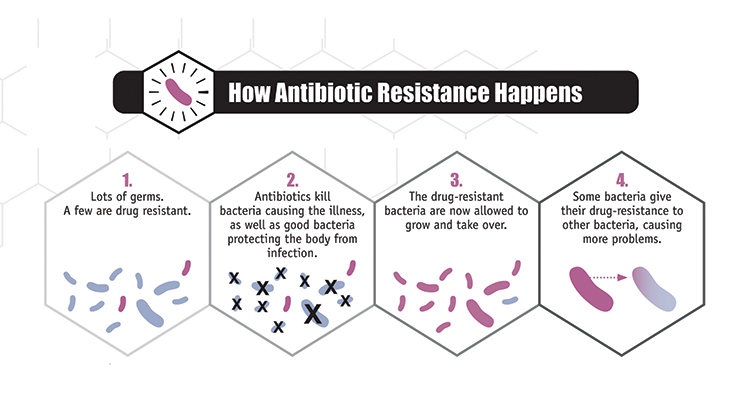
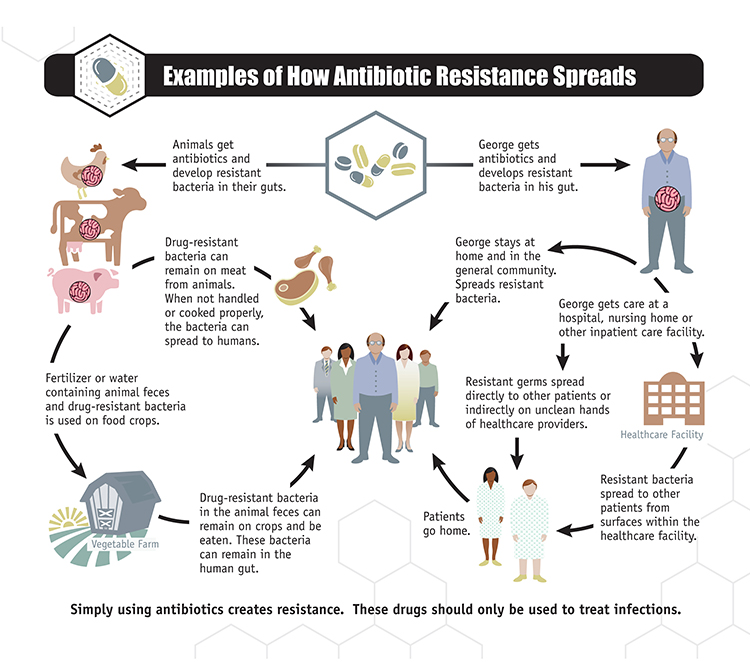




 I turned 60 on the 3rd of May. The year leading up to that was traumatic for me because, psychologically, it felt like a turning point in life – a long and enjoyable career behind me but a rapidly foreshortening window ahead! Surprisingly, things suddenly looked up when the birthday actually arrived (last Tuesday) – I didn’t really feel any different!
I turned 60 on the 3rd of May. The year leading up to that was traumatic for me because, psychologically, it felt like a turning point in life – a long and enjoyable career behind me but a rapidly foreshortening window ahead! Surprisingly, things suddenly looked up when the birthday actually arrived (last Tuesday) – I didn’t really feel any different!

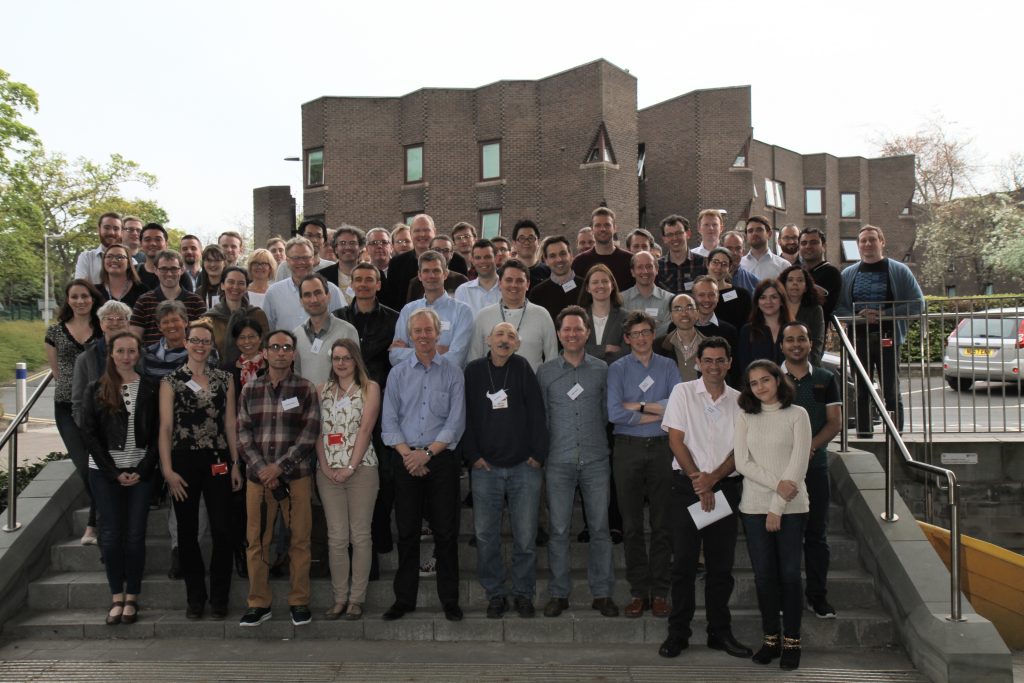
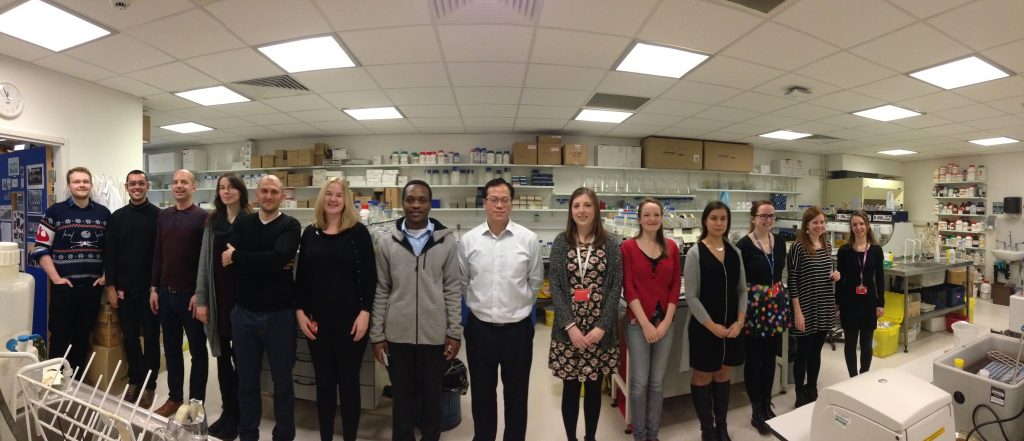
 We wondered how many of the past members of the lab would be able to make it, given that they were scattered to the four corners of the earth and we had no funds to cover their expenses. To my astonishment and delight, over 20 lab alumni agreed to come, and we were able to fill all of the 16 speaker slots with alumni from outside Newcastle.
We wondered how many of the past members of the lab would be able to make it, given that they were scattered to the four corners of the earth and we had no funds to cover their expenses. To my astonishment and delight, over 20 lab alumni agreed to come, and we were able to fill all of the 16 speaker slots with alumni from outside Newcastle.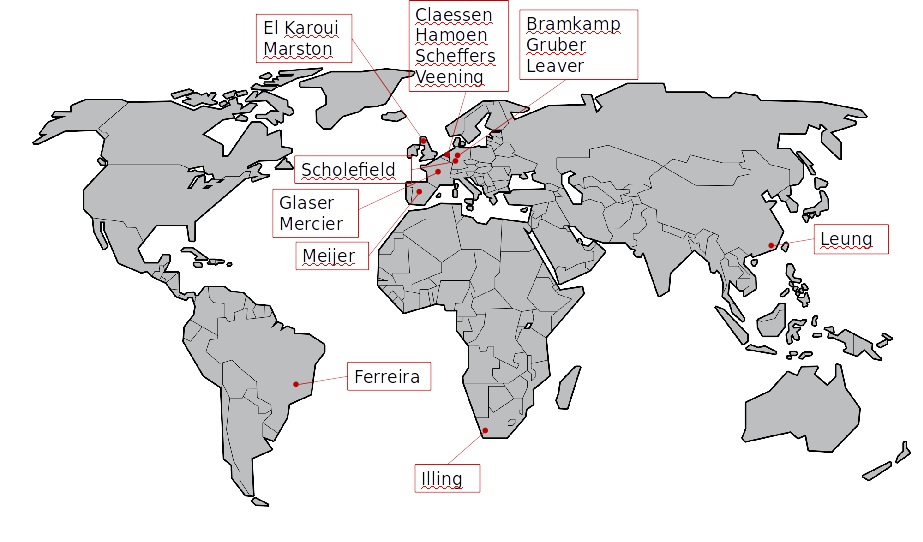




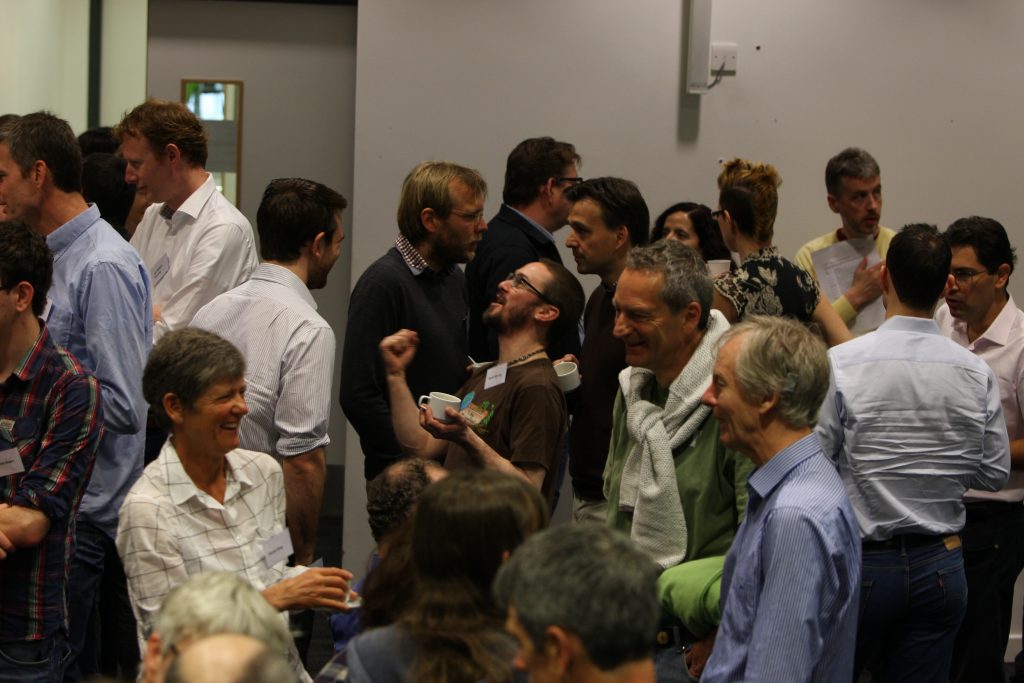
 A day of great science – and warm friendship
A day of great science – and warm friendship




 The only thing that I can think of is that if you can once assemble a critical mass of good scientists, which includes a few very good people, the culture or ecology of the lab becomes self sustaining. New people coming into the lab are well trained and inspired by the existing members, and they in turn go on to succeed in their own right. My job mainly becomes one of trying to make sure that the environment and infrastructure of the lab is capable of enabling good scientists to achieve their potential.
The only thing that I can think of is that if you can once assemble a critical mass of good scientists, which includes a few very good people, the culture or ecology of the lab becomes self sustaining. New people coming into the lab are well trained and inspired by the existing members, and they in turn go on to succeed in their own right. My job mainly becomes one of trying to make sure that the environment and infrastructure of the lab is capable of enabling good scientists to achieve their potential.









