In a recent issue of Science, the discovery of a key mechanism for bacterial cell division was reported. This work was carried out by Dr Seamus Holden’s lab (ICaMB) in collaboration with Professor Cees Dekker (TU Delft), Professor Yves Brun (Indiana University), Professor Mike VanNieuwenhze (Indiana University) and Professor Ethan Garner (Harvard University). Here, Seamus tells us about this discovery and what its implications could be for antimicrobial research.
In a recent issue of Science, the discovery of a key mechanism for bacterial cell division was reported. This work was carried out by Dr Seamus Holden’s lab (ICaMB) in collaboration with Professor Cees Dekker (TU Delft), Professor Yves Brun (Indiana University), Professor Mike VanNieuwenhze (Indiana University) and Professor Ethan Garner (Harvard University). Here, Seamus tells us about this discovery and what its implications could be for antimicrobial research.
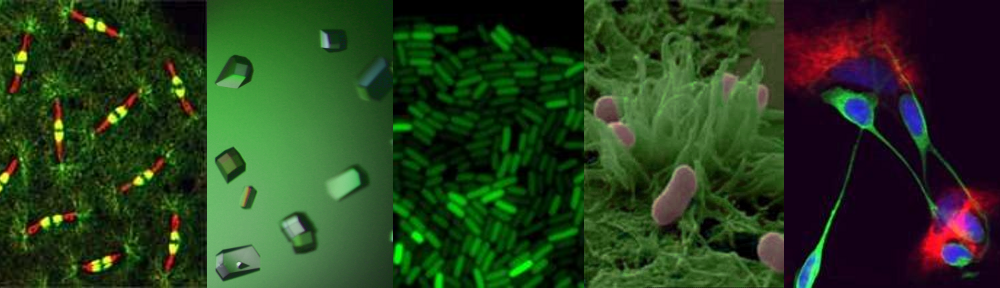
Bacterial cell division is a lovely mechanistic problem in biology: how do the simplest living organisms build a crosswall at mid cell, against very high outwards pressure (think of a racing bike tyre), without bursting? A ring of protein filaments forms around the future division site, and enzymes associated with this ring build a new crosswall that cleaves the bacteria in half. But what has remained completely mysterious is how these proteins work together as a single nanoscale machine to cut the bacterial balloon skin (cell wall) in two.
Working together with collaborators in Delft, Indiana and Harvard, we tracked the organization and motion of key division proteins as they build the dividing crosswall, and the organization of the newly built crosswall itself. We began by examining the motion of FtsZ, a cytoskeletal filament that is required for cell division – cytokinesis – in bacteria and is related to the tubulin cytoskeletal protein found in eukaryotic cells. Using high-resolution microscopy techniques, we found that FtsZ filaments move around the division site, traveling around the division ring. We imaged the motion of individual cell wall synthesis enzymes, and saw that the synthesis enzymes ride on FtsZ filaments, building new cell wall as they travel along the division site. This causes the cell wall to be synthesized in discrete sites that travel around the division site during cytokinesis, a process which we were able to observe directly by using dyes that label the bacterial cell wall. Using a variety of experimental techniques, we were able to speed up or slow down how fast FtsZ rotated around the cell. Strikingly, we found that the speed of FtsZ filament motion determines how fast the cell can divide. When FtsZ moves more rapidly, cell wall is produced more quickly, and cytokinesis happens faster. This shows that the motion of FtsZ is the critical overall controller of cell division.
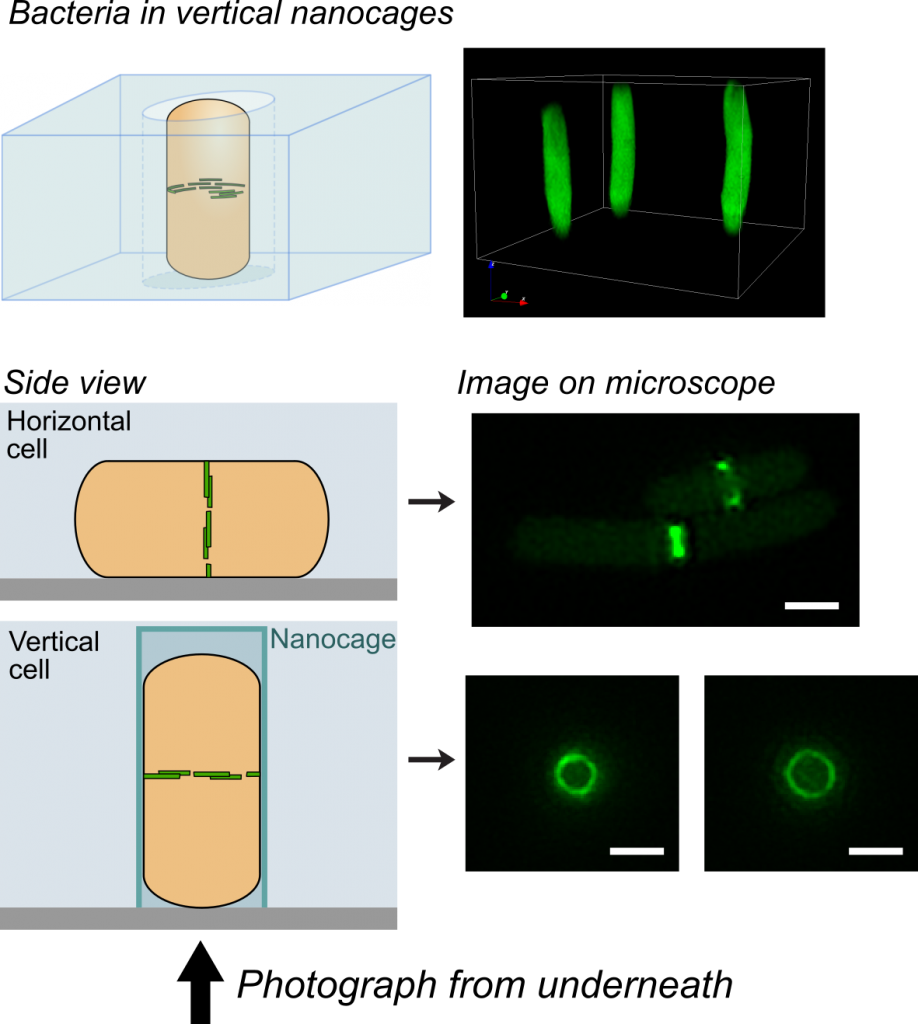
One challenge that we faced was trying to look at the division proteins in actively dividing cells. At the earliest stages of division, it was possible to image division protein organization because the proteins in the partially assembled ring are sparsely distributed. However, a new strategy was required to measure how the dense protein network of actively dividing cells was organized. Normally, bacteria are immobilized flat on a microscope slide, and imaged from underneath, but unfortunately this places the division ring side-on, obscuring the motion and organization of division proteins. To solve this problem, we used nanofabrication technology, originally developed to manufacture computer chips, to create tiny gel nanocages to trap bacteria in an upright position.
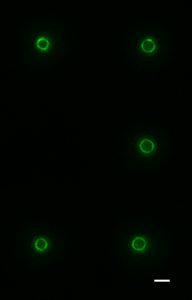
By trapping individual bacteria upright, we were able to rotate the cell division ring so that it was fully visible on our high resolution microscope. This revealed the dynamic motion of FtsZ filaments as they travel around the entire division site:
Together, these results revealed the basic mechanistic principles of bacterial cell division: that the building of the division crosswall is orchestrated by moving cytoskeletal filaments. Previously, the cytoskeleton was thought to serve as a static scaffold, recruiting other molecules and perhaps exerting some force to divide the cell. This new work demonstrates that all the components of cell division are in constant, controlled motion around the division site, driven by the fundamental dynamics of the cytoskeleton.
In the longer term, this study could open up novel antibiotic targets. Based on the discovery that the treadmilling motion of the bacterial cytoskeleton is critical for division, it may be possible to develop new drugs that specifically inhibit this motion, similar to how the chemotherapy drug taxol suppresses the motion of the cytoskeleton in cancer cells.
==================================
Links
Explanatory animation: https://youtu.be/6dq2_gqKPfU (Animation credit TU Delft / Scixel)
Nanocage Video: nanocage-movie-2.
Science report: http://science.sciencemag.org/content/355/6326/739
Press release: http://bit.ly/2kwsnJf